The human immune system is an astonishing diagnostic system, continuously adapting itself to detect any signal of disease in the body. Essentially, the state of the immune system tells a story about virtually everything affecting a person’s health. What if we could “read” this story? Our scientific understanding of human health would be fundamentally advanced. More importantly, this would provide a platform for a new generation of precise medical diagnostics and treatment options. We are partnering with Adaptive Biotechnologies (opens in new tab) to develop the machine learning and biotechnology tools that will allow us to realize this dream.
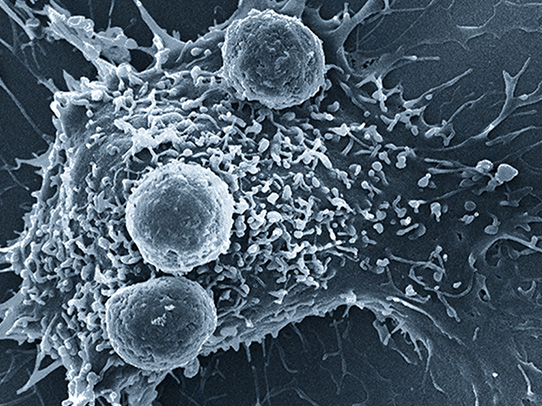
The Antigen Map Project
The Antigen Map is the complete mapping of which T-cells bind to which antigens. If we knew nature’s map, we could read the story about what your T-cells are currently targeting or have targeted in the past. Although the scale of the complete map is enormous, using a combination of high-throughput sequencing, functional assays and cloud-scale machine learning, we are learning to decode that map for disease-specific antigens. As this map comes into focus, we will unlock a new approach to diagnosing disease based on reading your immune system.

Blood sample
1 mL of blood contains about 1 million T-cells, each of which is genetically programmed to target a specific antigen your body is trying to control.

Immunosequencing
Adaptive’s immunoSEQ technology sequences the T cell receptor (TCR) gene that encodes each T-cell’s antigen specificity.

Machine learning
Microsoft is developing machine learning technologies that translate TCR sequences to the antigens they recognize.

Empowered care
Information about antigens your T-cells are actively targeting provides a basis for diagnosing cancers, infections and autoimmune diseases.
Learning to Decode the Immune System to Diagnose Disease
Adaptive published the original proof-of-concept for this project (Emerson, et al. Nat Genet. 2017 (opens in new tab)), demonstrating that shared TCRs can be used as a basis for diagnosing viral infection. Since then, we have further demonstrated the applicability of this work in covid, Lyme and other diseases (see publications). Despite the enormous complexity of the immune response, the elegant simplicity of this approach shows how common immunological solutions to pathogens can be used to provide a window into what the immune system is actively targeting. In principle, this approach can be extended and augmented with antigen-specific binding data to provide diagnostics for cancers, infections and autoimmune diseases. To date, we have sequenced the T-cell repertoires of tens of thousands of individuals affected by one of the several specific diseases we are initially pursuing based on unmet clinical need and our understanding of the underlying immunology:

Oncology
- Ovarian cancer
- Pancreatic cancer
Autoimmune Disease
- Type I diabetes
- Celiac disease
- Crohn’s disease
Infectious Disease
- COVID-19
- Lyme disease

Other diseases
- Other diseases with an unmet need for a blood-based diagnostic
More information on the antigen map project (opens in new tab) and the clinical development pipeline (opens in new tab) is available at Adaptive’s website.

